Origami is a craft that transforms the simplest materials into intricate works of art. Precise folds and careful planning draw fantastic creatures from an otherwise unremarkable sheet of paper: cranes, dragons, lilies, and…scaffolds for complex nanomachinery?
Through careful engineering and clever implementation of standard molecular techniques, scientists have found a way to fold long DNA helices into elaborate structures that they call DNA origami. This exciting technique—a very, very distant relative of the ancient art of paper folding—enables the creation of scaffolds for nanocomputers and nanorobotics that are mere tens of nanometers in length.
Fun Sized
To put the size of structures made with DNA origami into perspective, a human hair is on average 75,000 nanometers in diameter. Each seasonal flu virion has a diameter of only about 80–120 nanometers, and the individual proteins they use to invade to human cells are each only about 12 nanometers long. A nanometer is tiny.
Structures this small have become a Holy Grail of sorts for computer science and biomedicine, and have been proposed for use in a wide variety of applications, including drug delivery vehicles and switches for algorithmic computing. Advances in nanotechnology can help make many common materials lighter and stronger, electronics cheaper and more energy efficient, and medicines safer and easier to administer.
Nanoengineers are currently using DNA origami scaffolds as substrates for nanoelectric circuitry for devices that transfer information at the speed of light, while being much smaller than even the finest fiber optic cables. Researchers are even using DNA origami in their attempts to replicate photosystem II, which splits water into one oxygen and two hydrogen atoms during photosynthesis, to cheaply, safely, and efficiently turn water into hydrogen fuel.
Forming the Foundation
These DNA nanostructures could self-assemble into scaffolds that will hold the components of nanomachines in place with great precision.
Though some techniques have existed since the 1960s to try to make structures as small as those achieved by DNA origami from other materials, they are prohibitively labor-intensive. As a result, nanoengineers have searched for more efficient ways to create complex structures on a nanoscale.
Thus, Nadrian Seeman of SUNY Albany proposed the idea of using DNA as a framework for nanotechnology in 1982. His idea was that Watson-Crick pairing of the bases that make up DNA could be used for the rational design of tiny structures composed of DNA helices. This tendency of nucleobases to pair specifically with their complementary base (i.e., adenine to thymine and guanine to cytosine) allows scientists to design DNA sequences that are programmed to self-assemble into predetermined shapes.
The idea is quite elegant in its simplicity; why engineer building blocks to form these tiny structures when molecules already exist whose intrinsic properties allow us to do just that? These DNA nanostructures could self-assemble into scaffolds that would hold the components of nanorobots and nanocircuits in place with great precision, solving many current problems in nanoengineering. However, there were limitations to DNA nanoengineering that had to be solved before researchers could unleash their army of nanorobots on the world.
The Art of Invention
In the early days of DNA nanoengineering, researchers managed to design rudimentary self-assembling shapes and even simple machines like tweezers. However, they were limited by the technology of the time; manufacturers could only produce DNA strands of about 120 base pairs at most, severely limiting the complexity of the structures they could generate.
However, in 2006, Paul Rothemund of the California Institute of Technology came up with an idea that changed everything. Rather than try to manufacture specific sequences that would fold into a structure, Rothemund took a naturally occurring, very long strand of DNA – an entire viral genome, in fact – and ensured the strand of DNA stayed folded in the correct shape using what he called “staples.†These short strands of DNA were designed to use Watson-Crick pairing to bind the longer strand at specific sites, pulling the structure into the desired shape.
Trimming the Excess
Recent advances in DNA origami methods made DNA structures even more flexible and stable, and thus more useful for practical applications. Using principles based in graph theory, the branch of mathematics that studies how complex sets of points can be connected into a network, the Högberg group at Sweden's Karolinska Institutet created a simple, automated method to design hollow three-dimensional shapes, including a helix, a ball, and the Stanford bunny.
To make these complicated structures using as little DNA as possible, the Högberg lab designed a program that determines a route for folding a strand of DNA such that each edge of a structure consists of a single DNA helix, with the occasional “helper helix†doubling up on a side that needs a little more support. For example, to build a pyramid, rather than folding helices over each other until a solid pyramid has formed, the program folds the strand so that it draws just the edges of the shape, resulting in a hollow, balloon animal-like pyramid.
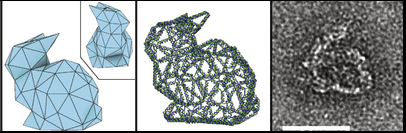

Reducing the amount of DNA needed to create nanostructures solves several major problems that had kept DNA origami from being used in practical applications in the past. If less DNA is needed for each edge of the shapes, the same number of DNA base pairs can create much more complex shapes with greater resolution.
Additionally, because very densely packed DNA requires ambient salt concentrations much higher than those found in a human body, medical applications were previously out of the question for DNA origami structures. The fact that less DNA is used to make Högberg's hollow three-dimensional scaffolds means that the buffers used to make folded DNA nanostructures don't need nearly as much salt to keep the DNA happy. In fact, Högberg and his colleagues successfully tested this new method in buffers commonly used in biomedical research that have salt concentrations similar to those found under physiological conditions.
Shaping the Future
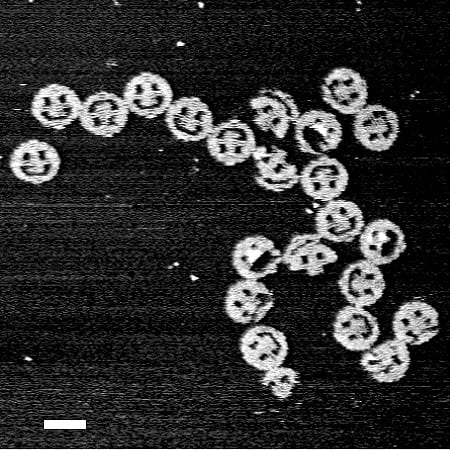
Though origami purists would have frowned upon the use of staples to create folded shapes, the scientific community has embraced DNA origami as a way to open the doors to an infinitesimal world of possibility. Rothemund and Högberg used DNA origami to fold DNA helices into microscopic smiley faces and bunnies, but with a bit of tweaking, it could be applied to building the complex nanomachines that scientists have spent decades striving for.
For more on the exciting world of nanotechnology, don't miss Graham Grable's podcast series on graphene and also carbon nanotubes.
About the Author
 Tara Bracken is a Ph.D. candidate at the University of Georgia Department of Infectious Diseases specializing in Global Health. Tara's research focuses on how the host's response to disease worsens malaria infection in children under the age of five and pregnant women. A true graduate student at heart, Tara is roughly 55% water and 45% caffeine. If you're a nerd and like fun, be sure to check out her blog, Of Microbes and Men or you can connect with her on Twitter at @microbesandmen! Tara Bracken is a Ph.D. candidate at the University of Georgia Department of Infectious Diseases specializing in Global Health. Tara's research focuses on how the host's response to disease worsens malaria infection in children under the age of five and pregnant women. A true graduate student at heart, Tara is roughly 55% water and 45% caffeine. If you're a nerd and like fun, be sure to check out her blog, Of Microbes and Men or you can connect with her on Twitter at @microbesandmen! |