What is a map? What does it REALLY represent? You might answer a map generally represents the relationships of locations. However, maps have evolved substantially in modern times. For example, modern genomics and cartography? While they might seem like different worlds, they actually share much in common. Both focused on a similar goal – spatial understanding.
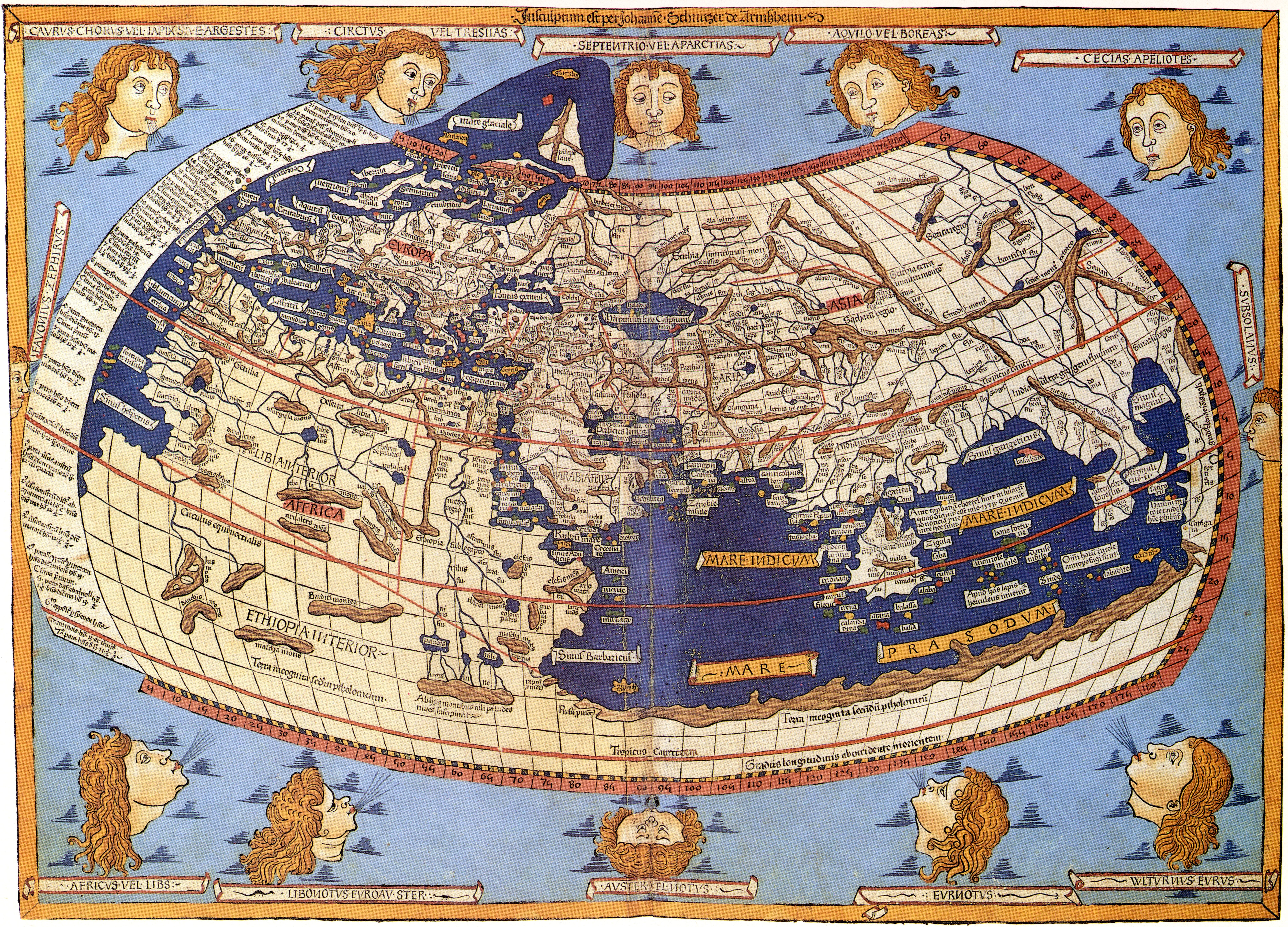
Image credit: Roderick Barron via Wikimedia Commons. Licensed under CC BY-NC 2.0.
But, before we dive into these similarities, it's important to understand the basics of what makes a map and how they altered human society. Broadly, we generate maps to understand spatial relationships. Over the centuries, this impacted human society in profound ways. Instead of individuals being limited in movement by locations they personally knew, humans exchanged information about the world and the surrounding landscape in the form of maps. This allowed knowledge to build on itself, allowing people to go farther into unexplored domains. We create genetic and genomic maps for this exact same reason. Scientists seek to make novel discoveries about the DNA in the human genome. So, by using a map, scientist are able to build off each others work, and make novel discoveries. So now that we understand why maps are so important, how do we make one for the human genome? How do we ensure that we as scientists aren't rediscovering the same locations on the map? And further, how do you know what to map?
So, imagine the human genome as a city. We have no idea where anything is, what anything does, and what locations in the city matter. Where do you start? Scientists are like explorers trying to understand this genomic city that already exists. So how do we get familiar with this new terrain? Well, a scientist would conduct experiments. These experiments are like sending out street view cars to take photos of the city. Each car takes photos, and notes locations that appear interesting and essential. Street signs, store fronts, neighborhoods, traffic lights (anything that seems important) – we take note. In the genome, these constitute things like genes, non-coding regions, or really repetitive DNA. All of this information pieced together forms the annotations on our genomic map.
As we gather information about this genomic city and find new features, we hope to gain a better sense about how the city works. For example we might start realizing that certain intersections are busier at certain times of the day, much like certain genes are turned on at very specific times. But, discovering more about the city isn't simply intriguing, mapping it provides us with valuable opportunities.
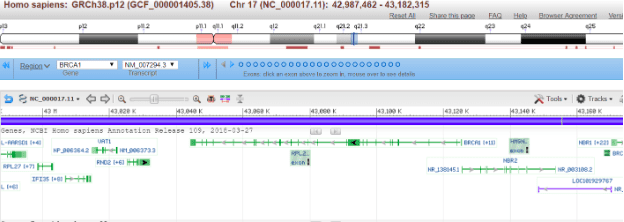
This maps power comes from a simple fact, it allows us to answer very specific questions about human biology. For instance, by finding variable regions between individuals, scientists can start to pinpoint what makes each of us unique. This invaluable approach applies to the realms of human health, helping us understand how our genetics contribute to diseases such as cancer. The genomic map completely revolutionized our understanding of what makes us us. But this map does not remain static.
Much like how early explorers refined the shape of our coastlines over decades of travel, our perception of the human genome evolves rapidly. Every time a scientist discovers something new about the genome (which is often) the map must be updated. The speed of these updates might take you by surprise. The initial map of the human genome was released to much fanfare in 2001, and less than a year later, we were already on version 11! With every update came incremental improvements, whether gene regions more accurately defined, or some unknown “gaps†being filled in. As it currently stands, the most recent version of the human reference genome is version number 38, with number 39 well on its way.
Even after 38 versions, we are still left with some massive outstanding questions: How many genes are in the genome? Where do the genes start and end? And what do most of these genes do? Much like Googleâ„¢ periodically sends out street view cars, we keep running more scientific experiments. One day we'll undoubtedly look back on our current version of the human genome, much like we look back on the ancient maps of Babylon, and realize that while it was a great start, there was still much unknown.
About the author:
 John Pablo Mendieta is a graduate student pursuing a PhD in bioinformatics and genomics at the University of Georgia. His specific interest lie at the intersection of agriculture, and genetic technologies. From Boulder Colorado, he enjoys the outdoors, science fiction, programming, and hip hop. You can email him at john.mendieta@uga.com or follow him on twitter @Pabster212.
John Pablo Mendieta is a graduate student pursuing a PhD in bioinformatics and genomics at the University of Georgia. His specific interest lie at the intersection of agriculture, and genetic technologies. From Boulder Colorado, he enjoys the outdoors, science fiction, programming, and hip hop. You can email him at john.mendieta@uga.com or follow him on twitter @Pabster212.